Research News and Highlights
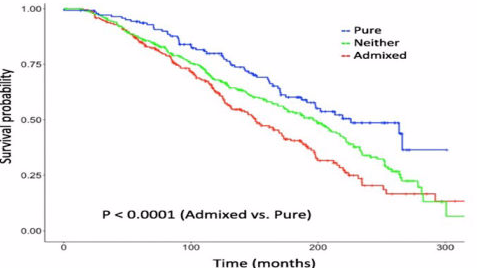
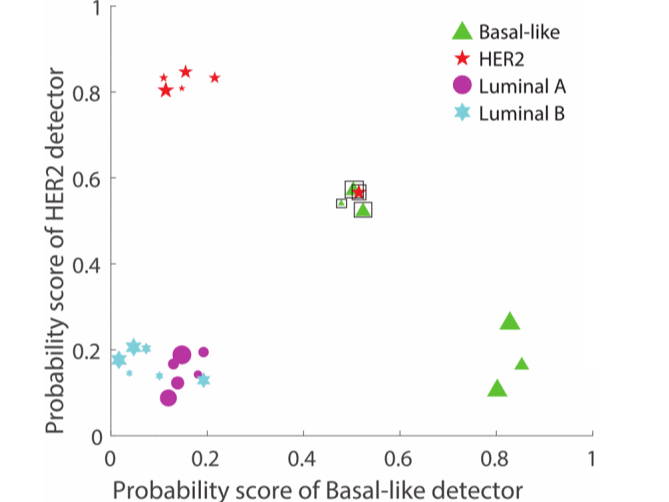
Breast cancers are subtyped and treated differently based on their molecular makeup. Even within a given subtype, some cancers are admixed with other subtypes. We proposed metrics to measure molecular heterogeneity in cancer and showed that these can predict surivival.
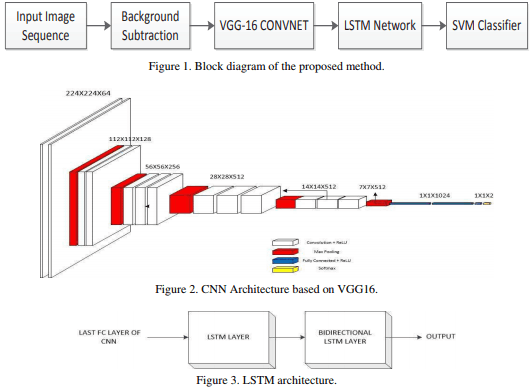
We organized an international competition to train deep neural networks to segment nuclei in digital pathology in MICCAI 2018, based on our previous work. Thirty six teams completed the challenge and their results make nucleus segmentation in H&E images almost a solved problem.
We were among the five finalists for the NVIDIA Global Impact Award in 2017 due to our work on computational pathology using machine learning and CNNs. In one of our projects, our team topped the automated HER2 IHC scoring challenge organized by TIA Lab at University of Warwick.
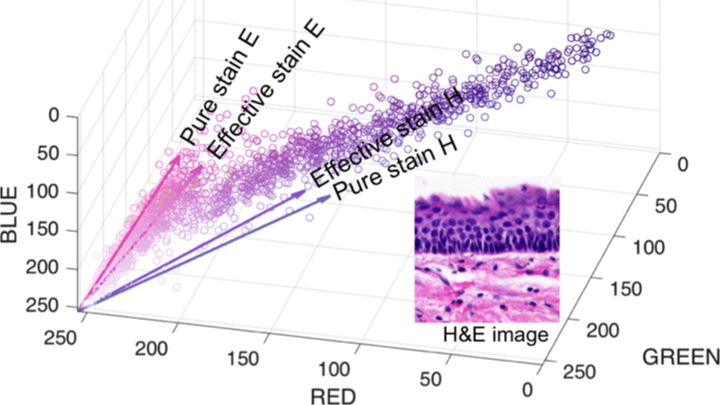
Our open source algorithm to color normalize histopathology slides has been widely used and cited due to its ability to preserve the underlying tissue structure while adapting images from a target domain to that of a source domain on which machine learning has been trained.